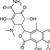
β-Apo-oxytetracycline, EvoPure (oxytetracycline-related compound E) is an oxytetracycline metabolite and impurity found in commercial oxytetracycline. It is formed via acid hydrolysis of oxytetracycline, but unlike other oxytetracycline metabolites, β-apo-oxytetracycline formation does not appear to be influenced by pH; however, it does form more rapidly with increasing temperature. β-Apo-oxytetracycline can be used as a reference standard during stability studies of oxytetracycline. It can be also be used to study the degradation pathway and products of tetracyclines. β-Apo-oxytetracycline, EvoPure is very slightly soluble in water.
We also offer:
- 4-Epi-oxytetracycline, EvoPure (oxytetracycline-related compound A)(O008)
- α-Apo-oxytetracycline, EvoPure (oxytetracycline-related compound D)(O009)
| Application | β-Apo-oxytetracycline, EvoPure is frequently used as a secondary standard. |
| Mechanism of Action | Oxytetracycline causes inhibition of protein synthesis. It binds to the 30S ribosomal subunit and prevents the amino-acyl tRNA from binding to the A site of the ribosome. |
| Spectrum | Broad-spectrum, including Gram-negative and Gram-positive bacteria. |
| Microbiology Applications | The fate of tetracyclines and their degradation products were studied under pH, chelation, and photodegradation. Studied whether they were potent to environmentally relevant sludge and soil bacteria. Tetracyclines photodecompose easily because they undergo direct photolysis and are converted to several products. MIC50 was 32 mg/L for tetracycline-sensitive strains of Pseudomonas, and > 32 mg/L for tetracycline-sensitive strains of Agrobacterium, Bacillus, E. coli. and Moraxella (Halling-Sorenson et al, 2002). |
| Molecular Formula | C22H22N2O8 |
| Solubility | Soluble in basic solutions. Very slightly soluble in water (0.2 mg/ml). |
| References |
Halling-Sorensen B, Sengelov G and Tjornelund J (2002) Toxicity of tetracyclines and tetracycline degradation products to environmentally relevant bacteria, including selected tetracycline-resistant bacteria. Arch. Environ. Contam. Toxicol. (2002) 42(3): 263-271 PMID 11910453 Lykkeberg AK, Halling-Sørensen B, Cornett C, Tjørnelund J and Honoré HS (2004) Quantitative analysis of Oxytetracycline and its impurities by LC-MS-MS. J. Pharm. Biomed. Anal. 34(2):325-332 PMID 15013146 Richeng X et al (2010) Hydrolysis and photolysis of Oxytetracycline in aqueous solution. J. Environ. Sci. and Health 45:73-81 PMID 20390934 |
| MIC | Bacillus cereus (ATCC 10702)| 1 - ?| 1302| Bacillus pumilus (ATCC 14884)| 2 - ?| 1302| Bacillus pumilus (ATCC 14884)| 2 - ?| 1183| Bacillus subtilis| 16 - ?| 1183| Bacillus subtilis| 16 - ?| 1183| Brochothrix thermosphacta| 124 - ?| 861| Burkholderia mallei (Pakistan)| ? - ?| 655| Clavibacter michiganensis subsp. michiganensis (185. 2. 1)| 4 - ?| 727| Clavibacter michiganensis subsp. michiganensis (C 8.2)| 4 - ?| 727| Clavibacter michiganensis subsp. michiganensis (IVIA-613)| 4 - ?| 727| Clavibacter michiganensis subsp. michiganensis (IVIA-873)| 4 - ?| 727| Diplococcus pneumoniae| 0.04 - 0.4| 1413| Enterococcus faecalis| 0.78 - >100| 1030| Enterococcus faecalis (broiler isolate)| 0.78 - >100| 1030| Enterococcus faecalis (cattle isolate)| 1.56 - >100| 1030| Enterococcus faecalis (pig isolate)| 0.78 - >100| 1030| Enterococcus faecium| 0.39 - >100| 1030| Enterococcus faecium (broiler isolate)| 0.39 - >100| 1030| Enterococcus faecium (cattle isolate)| 0.39 - >100| 1030| Enterococcus faecium (pig isolate)| 0.39 - >100| 1030| Enterococcus hirae| 0.39 - >100| 1030| Enterococcus sp. (Uraguay)| ≤0.25 - 2| 1391| Escherichia coli (ATCC 25922)| 0.25 - 2| 1127| Escherichia coli (K12 LE140)| 1.95 - ?| 1265| Escherichia coli (O157:H7)| 70 - ?| 861| Flavobacterium columnare| 15.63 - ?| 1347| Fusobacterium necrophorum| 0.03 - 4.1| 1444| Haemophilus influenzae| 1.6 - 6.3| 1413| Haemophilus parasuis (Spain)| 0.25 - 16| 768| Haemophilus parasuis (UK)| 0.25 - 16| 768| Klebsiella pneumonia (ATCC 10031)| 16 - ?| 1183| Listeria monocytogenes| 58 - ?| 861| Micrococcus kristinae| 1 - ?| 1302| Micrococcus kristinae| 4 - ?| 1183| Micrococcus luteus| 4 - ?| 1183| Mycoplasma bovis (Britain)| 1 - 128| 598| Mycoplasma bovis (Italy)| 0.12 - 4| 598| Mycoplasma bovis (Netherlands)| 8 - >64| 598| Mycoplasma gallisepticum| 0.05 - 200| 599| Mycoplasma gallisepticum (1975-1989)| 0.12 - 1| 606| Mycoplasma gallisepticum (1990-2000)| 0.05 - 200| 606| Mycoplasma gallisepticum (41-91)| 0.125 - ?| 595| Mycoplasma gallisepticum (ATCC 15302)| 0.125 - ?| 595| Mycoplasma hyopneumoniae| 0.25 - 4| 760| Mycoplasma hyopneumoniae| 0.12 - 2| 761| Mycoplasma hyopneumoniae (ATCC 25634)| 0.12 - ?| 594| Mycoplasma hyopneumoniae (ATCC 25634)| 0.12 - ?| 594| Mycoplasma hyopneumoniae (ATCC 25634)| 1 - ?| 594| Mycoplasma hyopneumoniae (field isolate + final)| 0.12 - >2| 761| Mycoplasma hyopneumoniae (field isolate + initial)| 0.03 - 2| 761| Mycoplasma hyopneumoniae (Final reading + Belgian)| 0.12 - >2| 594| Mycoplasma hyopneumoniae (Initial reading + Belgian)| 0.03 - 2| 594| Mycoplasma hyopneumoniae (J-strain + final)| 1 - ?| 761| Mycoplasma hyopneumoniae (J-strain + initial)| 0.12 - ?| 761| Mycoplasma hyosynoviae| 0.5 - >4| 1513| Mycoplasma iowae (1990-2000)| 0.025 - 100| 606| Mycoplasma iowae (90168)| 0.25 - ?| 595| Mycoplasma iowae (I 695)| 0.25 - ?| 595| Mycoplasma meleagridis (1975-1989)| 0.3 - 5| 606| Mycoplasma meleagridis (1990-2000)| 0.05 - 25| 606| Mycoplasma synoviae| 0.025 - 100| 599| Mycoplasma synoviae (1975-1989)| 0.06 - 0.08| 606| Mycoplasma synoviae (1990-2000)| 0.025 - 100| 606| Mycoplasma synoviae (WVU 1853)| 0.125 - ?| 595| Nocardia asteroides| 25 - 100| 1404| Paenibacillus larvae| 0.016 - 0.031| 1353| Pasteurella haemolytica| >64 - ?| 1022| Proteus vulgaris| 64 - ?| 1183| Proteus vulgaris (ATCC 6830)| 16 - ?| 1302| Proteus vulgaris (CSIR 0030)| 32 - ?| 1183| Pseudomonas aeruginosa (ATCC 19582)| 8 - ?| 1302| Pseudomonas flourescens| 40 - ?| 861| Salmonella enterica| 112 - ?| 861| Salmonella spp.| 2 - ?| 1302| Shigella flexneri| 1 - ?| 1302| Staphylococcus (coagulase-negative + Uruguay)| ≤0.25 - >64| 1391| Staphylococcus aureus| 76 - ?| 861| Staphylococcus aureus (ATCC 6538)| 4 - ?| 1183| Staphylococcus aureus (ATCC 6538)| 4 - ?| 1302| Staphylococcus aureus (OKOH1)| 1 - ?| 1302| Staphylococcus aureus (Uruguay)| ≤0.25 - >64| 1391| Staphylococcus epidermidis| 4 - ?| 1183| Streptococcus agalactiae (Uruguay)| ≤0.25 - 2| 1391| Streptococcus dysgalactiae (Uruguay)| >4 - >64| 1391| Streptococcus faecalis (ATCC 29212)| 8 - ?| 1183| Streptococcus iniae| 10 - ?| 1341| Streptococcus pneumonia (ATCC 6303 + quinolone-susceptible)| 0.5 - ?| 437| Streptococcus pneumonia (ATCC 7257 + quinolone-resistant)| 0.125 - ?| 437| Streptococcus uberis (Uruguay)| ≤0.25 - 64| 1391| Treponema hyodysenteriae| 0.78 - 25| 1427| Vibrio cholerae| 56 - ?| 861| Vibrio harveyi| 0.78 - ?| 1428| |