Bacitracin F, EvoPure is a highly purified form of Bacitracin fraction F. The EvoPure line of Bacitracins are bioactive, non-toxic congeners that have not shown toxicity to cell lines in eukaryotic cell culture.
Bacitracin F, EvoPure can be used to study properties and characteristics of Bacitracin F separately from other Bacitracin compounds found in standard grade bacitracin. Bacitracin F, EvoPure can also be used as an analytical standard.
For all Bacitracin products, click here.
| Mechanism of Action | Bacitracin prevents phosphorylation of bactoprenol, a transport protein which carries peptidoglycan components outside the cell membrane. Without the active phosphorylated bactoprenol, peptidoglycan synthesis cannot be completed and the cell lyses. Resistance to Bacitracin is understood to involve two mechanisms: A protein transporter (BcrABC) which pumps bacitracin out of the cell after it has entered, and via another protein (BacA) which provides the active phosphorylated bactoprenol from a different synthetic pathway. |
| Spectrum | Bacitracin primarily targets the cell wall in members of the Gram-positive bacteria including Streptococcus pyogenes and Staphylococcus aureus. |
| Microbiology Applications | Bacitracin is a useful tool to differentiate between ß-hemolytic, group A Streptococci (Streptococcus pyogenes) and ß-hemolytic Streptocococci of other groups. Bacitracin can be used as a supplement in chocolate agar to facilitate the isolation of Haemophilus influenzae. Bacitracin can be used to study the regulatory network in B. subtilis. By systematically analyzing the Bacitracin stimulon, authors can pinpoint the loci induced by Bacitracin (Mascher et al 2003). |
| Plant Biology Applications | Tobacco hairy roots and cell suspensions were used in plant transformation studies to produce full length murine IgG1 monoclonal antibody. Bacitracin has been shown to prevent degradation of peptides and hormones in plant systems. Treatment with Bacitracin was not sufficient to prevent loss of antibody from the cultures, but improved the growth rates by up to 53%. (Sharp and Doran, 1999). |
| Eukaryotic Cell Culture Applications | Bacitracin can be used for gene selection in cell culture. TOKU-E tested the Selectivity Factor values for the different grades of Bacitracin, and found that range was 41- 350 CulturePure grade versus 2-126 for USP grade, for the cell lines tested (HEK293, NIH/3T3, 293T, CHO-k1, B16). |
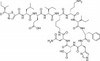
| Molecular Formula |
C66H98N16O17S |
| References |
Waterman et al. Webb et al. *** Bell, RG (1992) Preparative high-performance liquid chromatographic separation and isolation of Bacitracin components and their relationship to microbiological activity. J. Chromatog. 590:163-68 |
| MIC | Bacteroides fragilis (ATCC 25285)| >8 - ?| 650| Clostridium difficile| >256 - ?| 764| Clostridium perfringens (ATCC 13124)| 0.25 - ?| 650| Clostridium spiroforme (A7)| >8 - ?| 650| Clostridium spiroforme (A8)| >8 - ?| 650| Clostridium spiroforme (B61)| >8 - ?| 650| Clostridium spiroforme (BA 1)| >8 - ?| 650| Clostridium spiroforme (C121)| >8 - ?| 650| Clostridium spiroforme (CED 1)| >8 - ?| 650| Clostridium spiroforme (D11)| >8 - ?| 650| Clostridium spiroforme (D13)| 4 - ?| 650| Clostridium spiroforme (D14)| >8 - ?| 650| Clostridium spiroforme (NCTC 11493)| 8 - ?| 650| Clostridium spiroforme (RPO 980)| 8 - ?| 650| Diplococcus pneumoniae| 0.04 - 12.5| 1413| Enterococcus faecium| ≤1120 - ?| 837| Enterococcus faecium| ≤1120 - ?| 837| Escherichia coli| ≤560 - ?| 837| Escherichia coli| 800 - ?| 1541| Escherichia coli (14497)| 780 - ?| 1027| Escherichia coli (14497S)| 780 - ?| 1027| Escherichia coli (19568)| 780 - ?| 1027| Escherichia coli (19923)| 780 - ?| 1027| Escherichia coli (20384)| 780 - ?| 1027| Escherichia coli (20613)| 780 - ?| 1027| Escherichia coli (20619)| 780 - ?| 1027| Escherichia coli (3030-2)| 780 - ?| 1027| Escherichia coli (8829)| 780 - ?| 1027| Escherichia coli (8829S)| 780 - ?| 1027| Escherichia coli (ATCC 25922)| >8 - ?| 650| Escherichia coli (G58-1)| 1560 - ?| 1027| Escherichia coli (G58-1/pBR322)| 1560 - ?| 1027| Escherichia coli (JM109)| 390 - ?| 1027| Escherichia coli (JM109/pBR322)| 390 - ?| 1027| Fusarium oxysporum| 55 - ?| 1541| Fusobacterium necrophorum| 9.4 - 100| 1444| Haemophilus influenzae| >16 - ?| 120| Haemophilus influenzae| 50 - 400| 1413| Listeria innocua (7)| 50 - ?| 1024| Listeria monocytogenes (FBUNT)| 32 - ?| 1024| Listeria spp. (National Antimicrobial Resistance Monitoring Syste)| ≤8 - >128| 848| Neisseria gonorrhoeae| 800 - ?| 1541| Penicilium oxallicum| 50 - ?| 1541| Propionibacterium freudenreichii subsp. shermanii (131)| <1 - ?| 1029| Propionibacterium freudenreichii subsp. shermanii (NCIMB 10585)| <1 - ?| 1029| Propionibacterium freudenreichii subsp. shermanii (NCIMB 5959)| <1 - ?| 1029| Proteus vulgaris| 700 - ?| 1541| Pseudomonas aeruginosa| 700 - ?| 1541| Salmonella Agona (14749 + serovar B)| 3120 - ?| 1027| Salmonella Anatum (12453 + serovar E)| 3120 - ?| 1027| Salmonella Brandenburg (15865 + serovar B)| 3120 - ?| 1027| Salmonella Cerro (14294S + serovar D)| 1560 - ?| 1027| Salmonella Cerro (14824 + serovar D)| 1560 - ?| 1027| Salmonella Cerro (17390 + serovar D)| 1560 - ?| 1027| Salmonella Choleraesuis (12630 + serovar C1)| 3120 - ?| 1027| Salmonella Dublin (12133 + serovar D)| 3120 - ?| 1027| Salmonella Dublin (14178 + serovar D)| 1560 - ?| 1027| Salmonella Dublin (14178S + serovar D)| 1560 - ?| 1027| Salmonella Dublin (14294 + serovar D)| 1560 - ?| 1027| Salmonella Dublin (14347 + serovar D)| 1560 - ?| 1027| Salmonella Dublin (14534 + serovar D)| 1560 - ?| 1027| Salmonella Give (11520 + serovar E)| 3120 - ?| 1027| Salmonella Give (15065 + serovar E)| 3120 - ?| 1027| Salmonella Muenster (11814 + serovar E)| 3120 - ?| 1027| Salmonella Newport (12418 + serovar C2)| 3120 - ?| 1027| Salmonella Ohio (14497 + serovar C1)| 3120 - ?| 1027| Salmonella Stanley (17031 + serovar E)| 3120 - ?| 1027| Salmonella typhimurium| ≤1120 - ?| 837| Salmonella typhimurium (12948 + serovar B)| 3120 - ?| 1027| Staphylococcus aureus| 700 - ?| 1541| Staphylococcus aureus (8325)| 32 - ?| 806| Staphylococcus aureus (8325ΔpknB)| 32 - ?| 806| Staphylococcus aureus (ATCC 25923)| >8 - ?| 650| Staphylococcus aureus (ATCC 25923)| 16 - ?| 670| Staphylococcus aureus (erythromycin-resistant)| 2 - >4| 592| Staphylococcus aureus (erythromycin-susceptible + mupirocin-susceptible + oxacillin-susceptible)| ≤0.03 - >4| 592| Staphylococcus aureus (H14-KV)| 1 - ?| 670| Staphylococcus aureus (H5-KV)| 2 - ?| 670| Staphylococcus aureus (JSB20013)| 64 - ?| 748| Staphylococcus aureus (KSA)| 16 - ?| 670| Staphylococcus aureus (KVR)| 2 - ?| 670| Staphylococcus aureus (KVR-S)| 2 - ?| 670| Staphylococcus aureus (KVR-SR)| 8 - ?| 670| Staphylococcus aureus (KVR-Y)| 2 - ?| 670| Staphylococcus aureus (methicillin-resistant + mupirocin-resistant)| 2 - >16| 120| Staphylococcus aureus (methicillin-resistant)| 2 - >16| 120| Staphylococcus aureus (methicillin-susceptible)| 4 - >16| 120| Staphylococcus aureus (mupirocin-resistant)| 2 - >4| 592| Staphylococcus aureus (N315)| 16 - ?| 670| Staphylococcus aureus (N315ex)| 16 - ?| 670| Staphylococcus aureus (oxacillin-resistant)| 2 - >4| 592| Staphylococcus aureus (RN4220/tet(A)/pYH4)| 32 - ?| 748| Staphylococcus aureus (ΔIP)| 16 - ?| 670| Staphylococcus aureus (ΔIPH14)| 16 - ?| 670| Staphylococcus aureus (ΔIPH5)| 16 - ?| 670| Staphylococcus aureus (ΔIP-KV)| 2 - ?| 670| Staphylococcus auricularis (coagulase-negative + mupirocin-resistant)| 0.25 - >4| 592| Staphylococcus auricularis (coagulase-negative + mupirocin-susceptible + oxacillin-susceptible)| 4 - >4| 592| Staphylococcus auricularis (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus auricularis (LQC 5186)| 128 - ?| 836| Staphylococcus capitis (coagulase-negative + mupirocin-resistant)| 0.25 - >4| 592| Staphylococcus capitis (coagulase-negative + mupirocin-susceptible + oxacillin-susceptible)| 4 - >4| 592| Staphylococcus capitis (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus capitis (LQC 5184)| >128 - ?| 836| Staphylococcus caprae (LQC 5091)| 16 - ?| 836| Staphylococcus cohnii subsp. cohnii (LQC 5112)| 128 - ?| 836| Staphylococcus cohnii subsp. urealyticum (LQC 5195)| >128 - ?| 836| Staphylococcus epidermidis| 8 - >16| 120| Staphylococcus epidermidis (coagulase-negative + mupirocin-resistant)| 0.25 - >4| 592| Staphylococcus epidermidis (coagulase-negative + mupirocin-susceptible + oxacillin-susceptible)| 4 - >4| 592| Staphylococcus epidermidis (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus haemolyticus (coagulase-negative + mupirocin-resistant)| 0.25 - >4| 592| Staphylococcus haemolyticus (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus haemolyticus (LQC 5085)| >128 - ?| 836| Staphylococcus hominis (coagulase-negative + mupirocin-resistant)| 0.25 - >4| 592| Staphylococcus hominis (coagulase-negative + mupirocin-susceptible + oxacillin-susceptible)| 4 - >4| 592| Staphylococcus hominis (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus hominis (LQC 5012)| 16 - ?| 836| Staphylococcus intermedius (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus intermedius (LQC 5023)| 128 - ?| 836| Staphylococcus lugdunensis (coagulase-negative + mupirocin-susceptible + oxacillin-susceptible)| 4 - >4| 592| Staphylococcus lugdunensis (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus pyogenes (erythromycin-resistant)| 0.25 - >4| 592| Staphylococcus saprophyticus| 8 - >16| 120| Staphylococcus saprophyticus (coagulase-negative + mupirocin-resistant)| 0.25 - >4| 592| Staphylococcus saprophyticus (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus saprophyticus (LQC 5120)| >128 - ?| 836| Staphylococcus sciuri (LQC 5175)| 16 - ?| 836| Staphylococcus simulans (coagulase-negative + mupirocin-susceptible + oxacillin-susceptible)| 4 - >4| 592| Staphylococcus simulans (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus simulans (LQC 5187)| 128 - ?| 836| Staphylococcus warneri (coagulase-negative + mupirocin-resistant)| 0.25 - >4| 592| Staphylococcus warneri (coagulase-negative + mupirocin-susceptible + oxacillin-susceptible)| 4 - >4| 592| Staphylococcus warneri (coagulase-negative + oxacillin-resistant)| 0.25 - >4| 592| Staphylococcus warneri (LQC 5056)| >128 - ?| 836| Staphylococcus xylosus (LQC 5156)| >128 - ?| 836| Streptococci (Viridans group)| 2 - >16| 120| Streptococcus agalactiae| 2 - >16| 120| Streptococcus faecium| 850 - ?| 1541| Streptococcus pneumonia| 0.06 - >16| 120| Streptococcus pyogenes| 0.5 - >16| 120| Treponema hyodysenteriae| 12.5 - 50| 1427| |