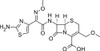
Cefpodoxime Sodium is a broad-spectrum, semi-synthetic, third-generation cephalosporin β-lactam and bactericidal agent that interferes with the bacterial cell wall. It is effective against a wide range of Gram-positive and Gram-negative bacteria. In vivo, it is de-esterified to the free acid and active metabolite, Cefpodoxime. Cefpodoxime, patented in 1980, is stable in the presence of β-lactamase enzymes, so many organisms resistant to penicillins and cephalosporins due to their production of β-lactamase, may be susceptible to Cefpodoxime. The compound is inactivated by certain extended-spectrum β-lactamases.
Cefpodoxime Sodium is soluble in DMSO.
We also offer:
| Mechanism of Action | Like β-lactams, cephalosporins interfere with PBP (penicillin binding protein) activity involved in the final phase of peptidoglycan synthesis. PBP’s are enzymes which catalyze a pentaglycine crosslink between alanine and lysine residues providing additional strength to the cell wall. Without a pentaglycine crosslink, the integrity of the cell wall is severely compromised and ultimately leads to cell lysis and death. Resistance to cephalosporins is commonly due to cells containing plasmid-encoded β-lactamases. However, like many cephalosporins, cefpodoxime is stable in the presence of β-lactamases. |
| Spectrum | Cefpodoxime Sodium is a broad-spectrum antibiotic which targets a wide variety of Gram-positive and Gram-negative bacteria. Notable exceptions include Pseudomonas aeruginosa, Enterococcus, and Bacterioides fragilis. |
| Microbiology Applications | Cefpodoxime Sodium is commonly used in clinical in vitro microbiological antimicrobial susceptibility tests (panels, discs, and MIC strips) against Gram-positive and Gram-negative microbial isolates. Medical microbiologists use AST results to recommend antibiotic treatment options. Representative MIC values include:
In vitro kinetic modeling can be used to study the pharmacokinetic-pharmacodynamic modeling of the antibacterial activity of Cefpodoxime. This approach has more detailed information than the MIC about the time course of efficacy (Liu et al, 2005). The suprior activity of the third-generation cephalosporins against Enterobacteriaceae is being challenged by the increasing frequency of pathogens with β-lactamase-mediated resistance, such as AMpC B-lactamases, ESBLs and carbapenemases. |
| Molecular Formula | C15H16N5NaO6S2 |
| References |
Alm RA, Johnstone MR and Lahiri SD (2015) Characterization of Escherichia coli NDM isolates with decreased susceptibility to aztreonam/avibactam: Role of a novel insertion in PBP3. J. Antimicrob. Chemother. 70(5):1420-1428 PMID 25634992 Doan VP et al (2019) Levofloxacin versus Cefpodoxime for antibacterial prophylaxis in allogeneic stem cell transplantation. Biol. Blood Marrow Transplant. 25(8):1637-1641 Georgopapadakou NH (1992) Mechanisms of action of cephalosporin 3'-quinolone esters, carbamates, and tertiary amines in Escherichia coli. Antimicrob. Agents Chemother. 37(3):559-565 Lahiri SD, Giacobbe RA, Johnstone MR and Alm RA (2014) Activity of avibactam against Enterobacter cloacae producing an extended-spectrum class C β-lactamase enzyme. J. Antimicrob. Chemother. 69(11):2942–2946 PMID 24986496 Liu P, Rand KH, Obermann B and Derendorf H (2005) Pharmacokinetic-pharmacodynamic modelling of antibacterial activity of Cefpodoxime and cefixime in in vitro kinetic models. Int. J. Antimicrob. Agents 25(2):120-129 PMID 15664481 Sheppard, M, King A and Phillips I (1991) In vitro activity of Cefpodoxime, a new oral cephalosporin, compared with that of nine other antimicrobial agents. Eur. J. Clin. Microbiol. Infect. Dis. 10:573–581 Wise R, Andrews JM, Ashby JP and Thornber D (1990) The in-vitro activity of Cefpodoxime: A comparison with other oral cephalosporins. J. Antimicrob. Chemother. 25(4):541–550 PMID 2351624 Product References Cefpodoxime (TOKU-E) was used against Escherichia coli NDM isolates in microdilution MIC assays in: "Characterization of Escherichia coli NDM isolates with decreased susceptibility to aztreonam/avibactam: Role of a novel insertion in PBP3." (Alm et al, 2015) Cefpodoxime (TOKU-E) was used as a reference compound when characterizing the extended-spectrum AmpC (ESAC) β-lactamase enzymes in "Activity of avibactam against Enterobacter cloacae producing an extended-spectrum class C β-lactamase enzyme" (Lahiri et al, 2014) |
| MIC | Bacteroides bivius| 0.25 - >64| 1492| Bacteroides disiens| 0.25 - >64| 1492| Bacteroides distasonis| 0.25 - >64| 1492| Bacteroides fragilis| 0.25 - >64| 1492| Bacteroides fragilis| 8 - 128| 401| Bacteroides fragilis (NCTC 9343)| 32 - ?| 401| Bacteroides ovatus| 0.25 - >64| 1492| Bacteroides thetaiotaomicron| 0.25 - >64| 1492| Bacteroides vulgatus| 0.25 - >64| 1492| Brucella abortus| 0.25 - ?| 1492| Brucella melitensis| 0.25 - ?| 1492| Brucella suis| 0.25 - ?| 1492| Burkholderia cepacia| 16 - >128| 600| Citrobacter amalonaticus| 16 - ?| 1481| Citrobacter amalonaticus (4026 + β-lactamase positive)| 8 - ?| 1481| Citrobacter braakii| >16 - ?| 1481| Citrobacter farmeri| 1 - >16| 1481| Citrobacter freundii| 0.5 - >128| 600| Citrobacter freundii| 4 - >16| 1481| Citrobacter freundii (3757 + β-lactamase positive)| 128 - ?| 1481| Citrobacter spp.| 16 - >16| 1481| Citrobacter werkmanii| >16 - ?| 1481| Clostridium difficile| 64 - >64| 1492| Clostridium histolyticum| 0.25 - 16| 1492| Clostridium sordellii| 0.25 - 16| 1492| Clostridium sporogenes| 0.25 - 16| 1492| Enterobacter aerogenes| 0.25 - >64| 1492| Enterobacter aerogenes| 1 - >16| 1481| Enterobacter aerogenes (3701 + β-lactamase positive)| 32 - ?| 1481| Enterobacter amnigenus| >16 - ?| 1481| Enterobacter asburiae| >16 - ?| 1481| Enterobacter cloacae| 0.03 - >64| 1492| Enterobacter cloacae| 0.125 - >128| 600| Enterobacter cloacae| 8 - >16| 1481| Enterobacter cloacae (22491)| 2 - ?| 386| Enterobacter cloacae (3146 + β-lactamase positive)| 64 - ?| 1481| Enterobacter cloacae (53 + β-lactamase positive)| 32 - ?| 1481| Enterobacter hormaechei| 8 - >16| 1481| Enterobacter intermedius| >16 - ?| 1481| Enterobacter sakazakii| 0.13 - 1| 1492| Enterobacter taylorae| >16 - ?| 1481| Enterobacteriaceae| 0.06 - 128| 401| Enterococci| 1 - 128| 401| Enterococcus faecalis| 1 - >128| 600| Enterococcus faecalis| >64 - ?| 1492| Enterococcus faecalis (ATCC 29212)| >32 - ?| 401| Enterococcus faecium| >64 - ?| 1492| Enterococcus faecium| 128 - >128| 600| Enterococcus liquefaciens| >64 - ?| 1492| Escherichia coli| 0.125 - 8| 600| Escherichia coli| 0.12 - 512| 946| Escherichia coli| ? - ?| 946| Escherichia coli (ampicillin-resistant + ceftazidime-resistant)| 16 - 32| 1492| Escherichia coli (ampicillin-resistant + ceftazidime-susceptible)| 0.13 - 0.25| 1492| Escherichia coli (ampicillin-susceptible)| 0.13 - 0.5| 1492| Escherichia coli (ATCC 25922)| 0.25 - ?| 401| Escherichia coli (ATCC 25922)| 0.5 - ?| 290| Escherichia coli (ATCC 35218)| 0.5 - ?| 290| Escherichia coli (NCTC 10418)| 0.25 - ?| 401| Escherichia coli (OC4075)| 1 - ?| 290| Escherichia coli (OC4087)| >128 - ?| 290| Escherichia coli (OC4136)| >128 - ?| 290| Escherichia coli (OC4138)| 32 - ?| 290| Escherichia coli (OC4227)| 8 - ?| 290| Escherichia coli (OC4229)| 16 - ?| 290| Escherichia coli (OC4249)| 64 - ?| 290| Escherichia coli (OC6028)| 64 - ?| 290| Escherichia coli (OC6042)| 128 - ?| 290| Escherichia coli (OC6043)| >128 - ?| 290| Escherichia coli (OC6044)| >128 - ?| 290| Haemolytic streptococci| 0.015 - 0.12| 401| Haemophilus influenzae| ≤0.03 - 1| 500| Haemophilus influenzae| 0.032 - 0.5| 600| Haemophilus influenzae (ampicillin-resistant)| 0.03 - 1| 1492| Haemophilus influenzae (ampicillin-susceptible)| 0.03 - 0.5| 1492| Haemophilus influenzae (ATCC 49247)| 0.5 - ?| 401| Haemophilus influenzae (ESBL)| <0.03 - 0.12| 84| Haemophilus influenzae (ESBL)| ≤0.03 - 0.5| 114| Haemophilus influenzae (ESBL)| <0.03 - 0.5| 84| Haemophilus influenzae (ESBL)| ≤0.03 - 0.5| 195| Haemophilus influenzae (NCTC 11931)| 0.12 - ?| 401| Haemophilus influenzae (non-ESBL)| ≤0.03 - 0.5| 195| Haemophilus influenzae (non-ESBL)| ≤0.03 - 0.5| 114| Haemophilus parainfluenzae| 0.06 - 0.13| 1492| Haemophilus spp.| 0.06 - 0.5| 401| Hafnia alvei| 16 - ?| 1481| Helicobacter pylori| 0.5 - 4| 1492| Klebsiella oxytoca (ceftazidime-resistant)| 2 - ?| 1492| Klebsiella oxytoca (ceftazidime-susceptible)| 0.06 - 2| 1492| Klebsiella pneumonia| 0.063 - 32| 600| Klebsiella pneumonia (ATCC 13883)| ≤0.12 - ?| 290| Klebsiella pneumonia (ATCC 700603)| 8 - ?| 290| Klebsiella pneumonia (OC4074)| 128 - ?| 290| Klebsiella pneumonia (OC4076)| 32 - ?| 290| Klebsiella pneumonia (OC4105)| 32 - ?| 290| Klebsiella pneumonia (OC4110)| >128 - ?| 290| Klebsiella pneumonia (OC4239)| 64 - ?| 290| Klebsiella pneumonia (OC4244)| >128 - ?| 290| Klebsiella pneumonia (OC4250)| >128 - ?| 290| Klebsiella pneumonia (OC5064)| >128 - ?| 290| Klebsiella pneumoniae (ceftazidime-resistant)| 8 - 64| 1492| Klebsiella pneumoniae (ceftazidime-susceptible)| 0.03 - 4| 1492| Klebsiella rhinoscleromatis| >16 - ?| 1481| Listeria innocua| 2 - >64| 1492| Listeria ivanovii| 2 - >64| 1492| Listeria monocytogenes| 2 - >64| 1492| Listeria seeligeri| 2 - >64| 1492| Listeria welshimeri| 2 - >64| 1492| Moraxella catarrhalis| 0.06 - 0.5| 1492| Moraxella catarrhalis| 0.06 - 2| 84| Moraxella catarrhalis| 0.06 - 2| 114| Moraxella catarrhalis| 0.06 - 4| 195| Moraxella catarrhalis| 0.063 - 2| 600| Moraxella catarrhalis| ? - ?| 403| Moraxella catarrhalis| ≤0.03 - >4| 500| Morganella morganii| 0.13 - 64| 1492| Morganella morganii| 0.125 - 64| 600| Morganella morganii| 8 - >16| 1481| Neisseria gonorrhoeae (penicillin-resistant)| 0.004 - 0.06| 1492| Neisseria gonorrhoeae (penicillin-susceptible)| 0.004 - 0.06| 1492| Neisseria spp.| 0.002 - 0.06| 401| Pneumococci| 0.03 - 4| 401| Proteus mirabilis| 0.06 - 0.13| 1492| Proteus mirabilis| 0.032 - 0.125| 600| Proteus mirabilis| <=0.25 - 4| 1481| Proteus mirabilis (3750 + β-lactamase positive)| 8 - ?| 1481| Proteus penneri| 16 - ?| 1481| Proteus penneri (1767 + β-lactamase positive)| 32 - ?| 1481| Proteus vulgaris| 0.06 - 0.5| 1492| Proteus vulgaris| 0.032 - 16| 600| Proteus vulgaris| <=0.25 - >16| 1481| Proteus vulgaris (1405 + β-lactamase positive)| 128 - ?| 1481| Proteus vulgaris (1699 + β-lactamase positive)| 128 - ?| 1481| Proteus vulgaris (1765 + β-lactamase positive)| >128 - ?| 1481| Proteus vulgaris (1781 + β-lactamase positive)| 1 - ?| 1481| Providencia rettgeri| ≤0.004 - 8| 600| Providencia rettgeri| ≤0.004 - 8| 600| Providencia rettgeri| 0.008 - 2| 1492| Providencia rettgeri| <=0.25 - 0.5| 1481| Providencia stuartii| 0.06 - 0.5| 1492| Providencia stuartii| <=0.25 - >16| 1481| Pseudomonas aeruginosa| >128 - ?| 600| Pseudomonas aeruginosa (ATCC 27853)| >128 - ?| 401| Pseudomonas aeruginosa (carbenicillin-resistant)| >64 - ?| 1492| Pseudomonas aeruginosa (carbenicillin-susceptible)| >64 - ?| 1492| Pseudomonas aeruginosa (NCTC 10662)| 128 - ?| 401| Pseudomonas cepacia| 4 - >64| 1492| Pseudomonas fluorescens| 8 - >64| 1492| Pseudomonas putida| 8 - >64| 1492| Pseudomonas spp.| 0.25 - 128| 401| Pseudomonas stutzeri| 8 - >64| 1492| Salmonella Agona| 0.13 - 0.5| 1492| Salmonella Brandenburg| 0.13 - 0.5| 1492| Salmonella Enteritidis| 0.13 - 0.5| 1492| Salmonella typhimurium| 0.13 - 0.5| 1492| Serratia fonticola| >16 - ?| 1481| Serratia liquefaciens| 0.25 - >64| 1492| Serratia marcescens| 0.5 - >128| 600| Serratia marcescens| 0.25 - >64| 1492| Serratia marcescens| 2 - >16| 1481| Serratia marcescens (167 + β-lactamase positive)| >128 - ?| 1481| Serratia marcescens (91 + β-lactamase positive)| >128 - ?| 1481| Serratia plymuthica| 8 - ?| 1481| Serratia rubidaea (3134 + β-lactamase positive)| 2 - ?| 1481| Serratia rubidaea (3139 + β-lactamase positive)| 4 - ?| 1481| Shigella boydii| 0.25 - 0.5| 1492| Shigella flexneri| 0.25 - 0.5| 1492| Shigella sonnei| 0.25 - 0.5| 1492| Staphylococci| 1 - 128| 401| Staphylococcus aureus (ATCC 25923)| 4 - ?| 401| Staphylococcus aureus (ATCC 29213)| 2 - ?| 401| Staphylococcus aureus (ATCC 6571)| 1 - ?| 401| Staphylococcus aureus (methicillin-resistant)| 16 - >64| 1492| Staphylococcus aureus (methicillin-resistant)| 1 - >128| 600| Staphylococcus aureus (oxacillin-susceptible)| ? - ?| 403| Staphylococcus epidermidis (methicillin-resistant)| 4 - >64| 1492| Staphylococcus epidermidis (methicillin-resistant)| 4 - >128| 600| Staphylococcus epidermidis (methicillin-susceptible)| 0.25 - 16| 600| Staphylococcus epidermidis (methicillin-susceptible)| 0.5 - 4| 1492| Staphylococcus haemolyticus (methicillin-resistant)| >64 - ?| 1492| Staphylococcus hominis| 0.5 - 2| 1492| Staphylococcus intermedius| ? - ?| 946| Staphylococcus pseudintermedius| 0.25 - 8| 1397| Staphylococcus simulans| 0.5 - 2| 1492| Staphylococcus warneri| 0.5 - 1| 1492| Stenotrophomonas maltophilia| >128 - ?| 600| Streptococci| ? - ?| 403| Streptococci (hemolytic + group A)| 0.008 - 0.5| 1492| Streptococci (hemolytic + group B)| 0.03 - 0.25| 1492| Streptococci (hemolytic + group C)| 0.015 - 0.03| 1492| Streptococci (hemolytic + group G)| 0.015 - 0.06| 1492| Streptococcus agalactiae| 0.032 - 0.063| 600| Streptococcus milleri (group)| 0.015 - 0.25| 1492| Streptococcus mitior| 0.015 - 0.13| 1492| Streptococcus pneumonia| ? - ?| 403| Streptococcus pneumonia (ATCC 49619)| 0.12 - ?| 401| Streptococcus pneumonia (ATCC 5303 parent)| 0.015 - ?| 287| Streptococcus pneumonia (ATCC 6303 parent)| 0.03 - ?| 287| Streptococcus pneumonia (pbp2x and pbp2b or pbp1a and pbp2x genes abnormal + penicillin)| 0.125 - 8| 121| Streptococcus pneumonia (pbp2x gene abnormal + penicillin-intermidiate)| 0.063 - 2| 121| Streptococcus pneumonia (pbpla, pbp2x, and pbp2b genes abnormal + penicillin-resistant)| 1 - 32| 121| Streptococcus pneumonia (penicillin-intermediate)| 0.05 - 2| 469| Streptococcus pneumonia (penicillin-intermediate)| ≤0.03 - >4| 114| Streptococcus pneumonia (penicillin-intermediate)| <0.03 - >4| 84| Streptococcus pneumonia (penicillin-intermediate)| ≤0.03 - >4| 195| Streptococcus pneumonia (penicillin-intermediate)| ? - ?| 418| Streptococcus pneumonia (penicillin-intermediate)| 0.12 - 4| 287| Streptococcus pneumonia (penicillin-resistant)| 0.063 - 16| 600| Streptococcus pneumonia (penicillin-resistant)| 1.861 - 8| 469| Streptococcus pneumonia (penicillin-resistant)| ≤0.03 - >4| 114| Streptococcus pneumonia (penicillin-resistant)| <0.03 - >4| 84| Streptococcus pneumonia (penicillin-resistant)| ≤0.03 - >4| 195| Streptococcus pneumonia (penicillin-susceptible)| ? - ?| 418| Streptococcus pneumonia (penicillin-susceptible)| ≤0.03 - 4| 114| Streptococcus pneumonia (penicillin-susceptible)| <0.03 - 4| 84| Streptococcus pneumonia (penicillin-susceptible)| ≤0.03 - 4| 195| Streptococcus pneumonia (penicillin-susceptible)| 0.016 - 0.5| 121| Streptococcus pneumonia (penicillin-susceptible)| 0.023 - 0.064| 469| Streptococcus pneumonia (penicillin-susceptible)| 0.008 - 1| 600| Streptococcus pneumonia (penicillin-susceptible)| 0.008 - 2| 287| Streptococcus pneumoniae (penicillin-susceptible)| 0.015 - 0.13| 1492| Streptococcus pyogenes| ≤0.004 - 2| 600| Xanthomonas maltophilia| >64 - ?| 1492| Yersinia enterocolitica| 0.5 - 2| 1492| |